Download the Powerpoint Presentation for Cryoelectron Microscopy Here
Abstract:
The integration of biological and physical sciences has been vital to the development of structural information for a number of biologically relevant targets.1 The use of techniques like X-ray crystallography, nuclear magnetic resonance and electron microscopy have consistently blossomed over the years. Biological macromolecules, however, are found in a variety of size, shape and intrinsic properties that make it difficult to use a single technique for elucidation of structural information for macromolecules.2 In particular, there are macromolecular assemblies that are too large to be routinely analyzed by either X-ray crystallography or NMR. Electron microscopy, may offer a solution towards analyzing large macromolecular assemblies in their physiological state at subatomic resolutions.3
The discovery of electrons by J. J. Thomson in 1897 and their particle-wave dual nature paved the way for the discovery of the electron microscope by Ernst Ruska in 1931.2 Electrons may either be scattered or transmitted when interacting with biological material thereby yielding either surface or internal structural information, respectively.4
However, an important advance was necessary before electron microscopy could be used for structural analysis. The necessity of high vacuum and the possibility of radiation damage implied that the biological macromolecules had to be protected well in their physiological state through freezing the specimen into a vitreous form by plunging it into liquid ethane.5 Thus, originated cryoelectron microscopy as a powerful tool for studying biological materials under physiological conditions.6
Cryoelectron microscopy involves viewing unaltered macromolecular assemblies by placing them on a grid and obtaining images by detecting electrons that transmit through the specimen.7 The number of techniques using cryoelectron microscopy is constantly increasing. These include single particle reconstruction, 2D crystals, cryoelectron tomography and freeze trapping.8
Single Particle Reconstruction
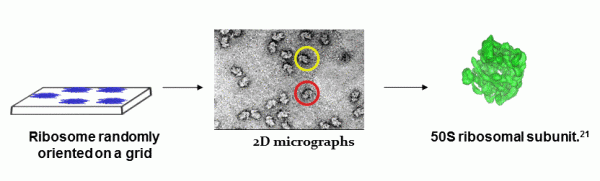
In principle, the technique involves the use of 2D micrographs to reconstruct a 3D model of the molecules imaged.9 An asymmetric, high molecular weight functional unit like the ribosome, which is difficult to visualize using traditional techniques like X-ray crystallography (due to difficulties in forming crystals and obtaining phase information), is easily observed in its native state by spreading a few microliters of a solution containing the ribosome onto a thin layer on a grid. This is followed by rapid freezing at vitreous conditions and imaging using an electron microscope. The numerous 2D images thus obtained are first processed to improve the signal to noise ratio followed by classification based on their orientation. Alignment and reconstruction are then achieved using computer programs resulting in the development of a 3D image of the ribosome in its native state (Figure 1).8,10,11 Important information on protein synthesis, ligand binding and RNA interaction can thus be obtained using this novel technique at medium resolutions of 7.5 to 25Å.9 Thus, utilization of this technique on similar asymmetric large particles has made it a useful tool in structural biology for viewing these complex macromolecules and studying their conformational changes at physiological conditions.
2D Crystals
Membrane-based proteins are an important class which include a number of receptors, transporters and channels.12 These are known to control a number of physiologically important functions in the body. However, structural information of these important drug targets embedded in lipid membrane is lacking due to technical difficulties in obtaining them in their natural state. Some of these proteins have an intrinsic property to form 2D crystals in the form of sheets or tubes.13 The ordered array of the 2D crystal structure makes it easier to assign angular information and obtain numerous images of the protein in a single orientation allowing accurate reconstruction of the 3D model at higher resolutions. Nicotinic receptors which form tubular 2D crystals were imaged using this technique and the 3D structure was obtained at resolution of 4Å.14 Further development of high throughput techniques to artificially obtain 2D crystals in a reconstituted lipid membrane have been valuable.15
Cryoelectron Tomography
Imaging equipment like magnetic resonance imaging (MRI) and computerized axial tomography (CAT) scan have been widely used in the past few decades to obtain 3D images of important organs in the human body. Similarly, a cryoelectron tomography can be used to obtain structural details of complex cellular organizations at subnanometer resolutions. The visualization of important cellular processes involved in large macromolecular assemblies like the Golgi apparatus, mitochondria, vesicles and microtubules can be achieved, giving a detailed understanding of how these complex systems function at a molecular level. Disease progression for a viral infection can also be studied by viewing the virus life cycle and the complex mechanism involved at various stages of the infection. The technique involves the use of cell cultures frozen under vitreous conditions, which are cryosectioned into thin slices of 1?m.16 The specimen is then viewed using a stage which is capable of tilting at angles of +70o.17 Radiation damage and inadequacies in sectioning techniques usually limit overall resolution obtained. However, the use of this informative technique has been increasing.
Freeze Trapping
Biologically relevant processes usually include intermediates which last for a few milliseconds. Structural information of these intermediates helps in gaining a thorough understanding of the mechanism of these processes. Chemical modification, use of faster techniques and freeze trapping are some of the techniques used to study these intermediates.18 Taking advantage of the low operating temperature of the cryoelectron microscope, the intermediate are freeze trapped and then viewed under an electron microscope. Nicotinic receptors were viewed in the closed state and later activated to the open state by spraying acetylcholine followed by rapid freezing and viewing under an electron microscope using a freefall device which is capable of studying reaction within timescales of 1 to 100 ms. Reconstructed 3D models were then used to study the detailed gating mechanism of these ligand gated ion channels.13,14 The mechanism thus obtained can then be used to identify new drug targets. Similarly, a wide variety of biological reactions like the light activated photocycle in bateriorhodopsin and the pH-induced fusion of viruses can also be studied using this technique.19,20
The applications thus indicate the versatility of cryoelectron microscopy as a powerful tool to gather important structural information for a variety of macromolecules, which include both easily purified homogenous molecules and complex heterogeneous systems. Limitations in resolution however, can be improved with the help of new advances in instrumentation and computer programs. Cryoelectron microscopy thus promises to continue to be an exciting tool in the field of structural studies and their interpretation for biologically relevant molecules.
References:
- Hasnain, S. S.; Wakatsuki, S. Biophysical methods: structure, function and dynamics studies of macromolecular assemblies using electrons, lasers, neutrons and X-rays. Curr. Opin. Struct. Biol. 2008, 18, 557-580.
- Campbell, I. D. The march of structural biology. Nat. Rev. Mol. Cell Biol. 2002, 3, 377-381.
- Jonic, S.; Venien-Bryan, C. Protein structure determination by electron cryo-microscopy. Curr. Opin. Pharmacol. 2009, 9, 636-642.
- Spahn, C. M. T.; Penczek, P. A. Exploring conformational modes of macromolecular assemblies by multiparticle cryo-EM. Curr. Opin. Struct. Biol. 2009, 19, 623-631.
- Taylor, K. A.; Glaeser, R. M. Retrospective on the early development of cryoelectron microscopy of macromolecules and a prospective on opportunities for the future. J. Struct. Biol. 2008, 163, 214-223.
- Zhou, Z. H. Towards atomic resolution structural determination by single particle cryo-electron microscopy. Curr. Opin. Struct. Biol. 2008, 18, 218-228.
- Henderson, R. Realizing the potential of electron cryo-microscopy. Q. Rev. Biophys. 2004, 37, 3-13.
- Frank, J. Single-particle reconstruction of biological macromolecules in electron microscopy – 30 years. Rev. Biophys. 2009, 42, 139-158.
- Llorca, O. Introduction to 3D reconstruction of macromolecules using single particle electron microscopy. Acta. Pharmacol. Sin. 2005, 26, 1153-1164.
- Frank, J. Cryo-electron microscopy as an investigative tool: the ribosome as an example. BioEssays. 2001, 23, 725-732.
- Gabashvili, I. S.; Agrawal, R. K.; Spahn, C. M. T.; Grassucci, R. A.; Svergun, D. I.; Frank, J.; Penczek, P. Solution structure of the E. coli 70S ribosome at 11.5 Å resolution. Cell 2000, 100, 537-549.
- Fujiyoshi, Y.; Unwin, N. Electron crystallography of proteins in membranes. Curr. Opin. Struct. Biol. 2008, 18, 587-592.
- Unwin, N. Acetylcholine receptor channel imaged in the open state. Nature 1995, 373, 37-43.
- Miyazawa, A.; Fujiyoshi, Y.; Unwin, N. Structure and gating mechanism of the acetylcholine receptor pore. Nature 2003, 423, 949-955.
- Renault, L.; Chou, H.; Chiu, P.; Hill, R. M.; Zeng, X.; Gipson, B.; Zhang, Z.; Cheng, A.; Unger, V.; Stahlberg, H. Milestones in electron crystallography. J. Comput. Aided Mol. Des. 2006, 20, 519-527.
- Koning, R. I.; Koster, A. J. Cryo-electron tomography in biology and medicine. Ann. Anat. 2009, 191, 427-445.
- Jonic, S.; Sorzano, C. O. S.; Boisset, N. Comparison of single-particle analysis and electron tomography approaches: an overview. J. Microsc. 2008, 232, 562-579.
- Moffat, K.; Henderson, R. Freeze trapping of reaction intermediates. Curr. Opin. Struct. Biol. 1995, 5, 656-663.
- Fuller, S. D.; Berriman J. A.; Butcher S. J.; Gowen B. E. Low pH induces swiveling of glycoprotein heterodimers in the Semliki forest virus spike complex. Cell 1995, 81, 715-725.
- Subramaniam, S.; Lindahl, M.; Bullough, P.; Faruqi, A. R.; Tittor, J.; Oesterhelt, D.; Brown, L.; Lanyi J.; Henderson, R. Protein conformational changes in the bateriorhodopsin photocycle. J. Mol. Biol. 1999, 287, 145-161.
- EMDB ID: 1019.