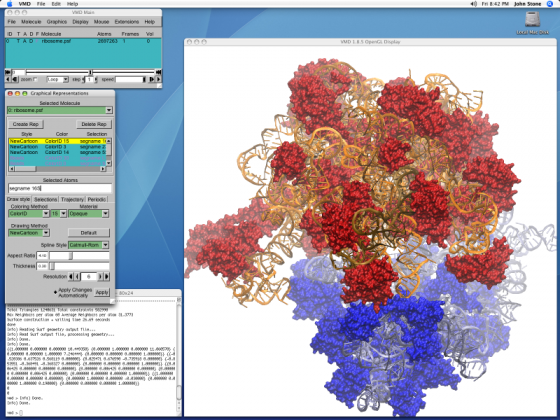
Visual Molecular Dynamics (or better known as VMD) is a software which has been developed by University of Illinois at Urbana Champaign, in order to visualize proteins, macromolecules, biological systems, nucleic acids, lipid bilayer assemblied etc. and also run molecular dynamics simulations on them.VMD is a computational chemistry or molecular modelling software that is free to be used for academic purposes and can be downloaded from their website. The rendering engine of VMD has been used widely to produce stunning images of proteins, and macromolecules and has been used on the cover pages of several textbooks and journals. VMD has been cited by over 4000 publications world wide.
Where Can You Get VMD?
Visual Molecular Dynamics (VMD) is free to be downloaded on the UIUC server. You will need to register (which is free) and login before you can download the software. The following page allows you to download VMD to work with Windows (XP/Vista/7), SolarisX86_64, MacOS X, Linux, Blue Waters, and AIX6_64. Thus it can be used across a wide variety of operating systems. It is also available in both 32-bit as well as 64-bit versions.
Download Page : http://www.ks.uiuc.edu/Development/Download/download.cgi?PackageName=VMD
Why Use VMD?
The advantage of using VMD over other visualization softwares is that VMD utilizes the graphics processing units (GPUs) of the computer to the fullest extent in order to render the large biomolecular complexes. By using the GPU, VMD can accelerate the calculations of electrostatic potential fields, visualization and analysis as well as modelling operations such as ion placement.
VMD can be used to read standard Protein Data Bank (PDB) files and display the contained structures. VMD also offers a wide variety of methods for rendering and coloring a molecules such as:
- simple points and lines
- CPK spheres and cylinders (ball and stick)
- licorice bonds
- backbone tubes and ribbons
- cartoon drawings
- and many more.
VMD can also be used to animate the structures and analyze the trajectory of a molecular dynamics (MD) simulation. VMD acts as a graphical user interface for external molecular dynamics programs by displaying and animating a molecule undergoing simulation on a remote computer. VMD can be interfaced with molecular dynamics simulation software such as NAMD (also developed by University of Illinois at Urbana Champaign) and SIGMA.
Sample Images, Movies and Media Generated Using VMD
VMD has been widely used to render images, media, science movies, posters and brochures to publish. Here is a link which will show you the sample media in the TCB Gallery:
http://www.ks.uiuc.edu/Gallery/
Scripts, Plugins and Add-Ons
VMD also has an extensive scripts library which provides a plethora of add-ons which you can put on your system to run various computational calculations. These include, but are not limited to, structure building scripts, analysis scripts, trajectory processing scripts, file conversion scripts, and many more. A programmer may even submit their own scripts and there are plenty of user-defined scripts also available.
Tutorials and Documentation
VMD has a very good documentation page which provides information on how the software works, as well as provides plenty of tutorials on running calculations. It also contains tutorials developed by the NIH Resource for Macromolecular Modeling and Bioinformatics. It even has tutorials for its integration with the molecular dynamics software such as NAMD. These tutorials are free to download and view.
Key Features of VMD
- Support for all major computer platforms
- Support for a variety of file formats including .pdb , .mol2 etc.
- Support for multicore processors
- Support for GPU accelerated computation
- Many excellent VMD tutorials developed locally, and by the research community at large
- No limits on the number of molecules, atoms, residues or number of trajectory frames, except available memory
- Many molecular rendering and coloring methods
- Stereo display capability
- Extensive atom selection syntax for choosing subsets of atoms for display (includes boolean operators, regular expressions, and more)
- Support for over 60 molecular file formats and data types through an extensive library of built-in file reader/writer plugins and translators
- VMD includes a multiple sequence alignment plugin, a unified bioinformatics analysis environment that allows one to organize, display, and analyze both sequence and structure data for proteins and nucleic acids.
- Ability to export displayed graphics to files which may be processed by a number of popular ray tracing and image rendering packages, including POV-Ray, Rayshade, Raster3D, and Tachyon.
- User-extensible graphical and text-based user interfaces, built-on standard Tcl/Tk and Python scripting languages
- Extensions to the Tcl language which enable researchers to write their own routines for molecular analysis
- Modular, extensible source code using an object-oriented design in C++, with a programmers guide describing the program architecture and source code
- Integration with the program NAMD, a fast, parallel, and scalable molecular dynamics program developed in conjunction with VMD in the Theoretical and Computational Biophysics Group at the University of Illinois. See the NAMD WWW home page for more info:
http://www.ks.uiuc.edu/Research/namd/ - VMD works well with projected display systems like the 3-D Projection Facility maintained by the Theoretical and Computational Biophysics Group.
- VMD can be used to concurrently display and interact with a running NAMD simulation.
Reference
- Humphrey, W., Dalke, A. and Schulten, K., “VMD – Visual Molecular Dynamics”, J. Molec. Graphics, 1996, vol. 14, pp. 33-38.
- http://www.ks.uiuc.edu/Research/vmd/

great